BAC Recombineering
Bacterial artificial chromosomes (BACs) are DNA vectors suitable for cloning and stably maintaining large DNA inserts of up to 300 Kb in E. coli. BACs can be genetically manipulated relatively easily and quickly to carry any desired modification using a homologous recombination-based genetic engineering technique known as BAC recombineering.
VectorBuilder offers a variety of BAC modification services, including placing reporters behind regulatory sequences on your BAC, introducing point mutations into genes of interest, transferring regions of the BAC onto a plasmid, and adding drug-selection or visualization markers to the BAC backbone. All you will need to do is send us a design request describing your experimental goal and we will take care of the rest for you starting from designing your BAC modification strategy and ordering your BAC to generating and validating your final modified BAC clones.
Click to view 1000+ citationsClick to view user testimonials
Service Details
Price and turnaround Price Match
The cost and the turnaround time for BAC recombineering projects are relatively high due to the amount of work involved. However, outsourcing to us is still far more economical and faster than doing it yourself. Due to the complexity and error-prone nature of the procedure, most people attempting it themselves can spend up to a year gathering reagents, learning the protocol, and troubleshooting before they succeed. You can instead spend your time on formulating hypotheses or designing experiments and leave the tedious cloning work to us. An overview of our available BAC recombineering services is shown in Table 1 and the breakdown of price and turnaround is shown in Table 2 below.
Table 1. Overview of BAC recombineering services.
| Available Services | Price (USD) | Turnaround | Note |
|---|---|---|---|
| Knockin | |||
| EGFP, mCherry, LacZ, GCaMP6s, Luciferase, DTR | $2,999 | 42-63 days | Drug-selection cassette used in one-step BAC modification is removed. |
| $4,098 | 63-84 days | Drug-selection cassette used in one-step BAC modification is removed; piggyBac ITRs are added to BAC backbone. | |
| Cre, CreERT2 | $7,179 | 98-119 days | Drug-selection cassette used in one-step BAC modification is removed. LoxP sites on BAC backbone are removed. piggyBac ITRs can be added to BAC backbone. |
| Custom DNA fragment | Please inquire. | ||
| Point mutation | |||
| Custom point mutation | $5,599 | 49-77 days | Two-step BAC modification. |
| $6,698 | 70-98 days | Two-step BAC modification; piggyBac ITRs are added to BAC backbone. | |
| Removal of loxP sites on BAC backbone | |||
| pBACe3.6 | $4,480 | 63-84 days | The loxP and loxP511 sites on pBACe3.6 backbone may interact with Cre to cause unwanted recombination in the genome. piggyBac ITRs can be added to BAC backbone. |
| Other BAC cloning vectors | Please inquire. | ||
| Transfer fragment from BAC to plasmid | |||
| pStart-K | $2,999 | 35-49 days | This price applies when the transferred fragment is under 30 kb. Higher prices may apply when the transferred fragment is over 30 kb. The pStart-K plasmid backbone can accommodate 60 kb insert (and possibly even longer). The pStart-K-ITR backbone contains PiggyBac ITRs that flank the insert, allowing PiggyBac-mediated genome integration of the insert. |
| pStart-K-ITR | |||
| Other plasmids | Please inquire. | ||
| Add visualization markers or other elements to BAC backbone* | |||
| Neo, Puro, piggyBac ITRs | $1,399 | 35-42 days | These elements are added to the loxP site on BAC backbone. |
| Other elements | Please inquire. | ||
*This service is available only for BACs that contain a loxP site in their backbones. Most BACs from the BACPAC Resource Center contain loxP sites.
Table 2. Breakdown of price and turnaround for BAC recombineering services.
| Service | Price (USD) | Turnaround |
|---|---|---|
| Design BAC modification strategy | Free | 1-4 days |
| Obtain BAC from vendor | $300 | 2 weeks |
| One-step BAC modification (without removing drug-selection cassette) | From $2,090* | 2-4 weeks |
| Optional: Remove drug-selection cassette used in one-step BAC modification | $350 | 1-2 weeks |
| Two-step BAC modification | From $5,099* | 4-8 weeks |
| Add drug-selection or visualization marker to BAC backbone** | From $1,099 | 1-2 weeks |
| Transfer region of BAC onto plasmid | $2,699 | 2-4 weeks |
| Validation of modified BAC | Free | 1 week |
*Price may vary depending on the complexity of the sequence being introduced.
**This service is available only for BACs that contain a loxP site on their backbones. Most BACs from the BACPAC Resource Center contain loxP sites.
Design BAC modification strategy
VectorBuilder’s experienced team of scientists will put together a detailed and highly effective strategy for achieving the BAC modification your need, all for free!
Obtain BAC from vendor
If you know the ID number of the BAC you need, just give us the number and we will buy it for you. If you don’t know which specific BAC to use, we will find the appropriate one for you based on the gene you want to study. The great majority of BACs are available from the BACPAC Resource Center at CHORI, but some BACs have to be obtained from other suppliers. This step is not needed if you send us your own BAC as a E. coli stock.
Validation of modified BAC
The modified region of the BAC will be confirmed by sequencing. Sometimes, BACs can undergo deletions due to the presence of repetitive sequences. To mitigate this possibility, we will re-transform the modified BAC to obtain a single clone and confirm that it has not undergone major deletions by performing PCR amplifications of multiple sites across the entire BAC.
Deliverables
The final deliverable for our BAC recombineering service is E. coli stock containing the modified and validated BAC. Multiple scales of high-quality BAC DNA can be purchased additionally: Miniprep (>10 ug), Midiprep (>100 ug), Maxiprep (>300 ug), Megaprep (>1 mg) and Gigaprep (>10 mg).
Customer-supplied materials
If customer-supplied BACs are needed, please submit your materials information on "Support" > "Materials Submission Form". Please strictly follow our guidelines to set up shipment to avoid any delay or damage of the materials. All customer-supplied materials undergo mandatory QC by VectorBuilder which may incur an additional QC charge starting from $120 per item. Please note that production cannot be initiated until customer-supplied materials pass QC.
Technical Information
Advantages and applications of BACs
While utilizing regular plasmid vectors is a standard and the most common approach for expressing transgenes of interest in cell culture and animals, the application of such vectors is sometimes limited by their ability to carry DNA sequences of only up to 30 Kb in length. BACs on the other hand, are suitable for cloning and stably maintaining large DNA inserts of up to 300 Kb in E. coli. BACs are based on the fertility (F) factor plasmid of E. coli which helps in the maintenance of one copy of the BAC per bacterial cell, thereby making them less susceptible to rearrangements and therefore, more stable compared to high copy number plasmids. Majority of BACs can be readily and inexpensively obtained from various bioresource centers such as the BACPAC Resource Center, the Arabidopsis Resource Center, The Sanger Institute and INRA BAC-YAC Resource Center as well as from certain commercial resources. These BACs can then be genetically manipulated relatively easily and quickly to carry any desired modification using BAC recombineering.
Due to their ability to confer stability to large pieces of DNA and the relative ease with which they can be engineered, BACs have been widely used in genome sequencing projects of several model organisms including human, mice and Arabidopsis thaliana. Other common applications of BACs include utilizing BAC library screening for positional cloning of disease-associated genes, creating transgenic cell lines or transgenic mice harboring BACs carrying wildtype, mutated or reporter tagged versions of genes or regulatory elements to study their roles in various disease and molecular pathways and generating infectious BAC clones carrying genomes of DNA or RNA viruses such as herpesvirus for studying infectious diseases caused by such viruses.
One-step BAC modification
This modification can be used to introduce either major sequence changes such as placing a reporter behind a gene’s promoter to track gene expression (Figure 1), or minor changes such as point mutations (Figure 2).
In the case of a major change (Figure 1), a DNA fragment containing your gene of interest (GOI) such as LacZ or GFP reporter, plus a drug-selection cassette flanked by FRT sites (along with flanking homology regions) is used to replace an endogenous region on the BAC via homologous recombination. The drug-selection cassette facilitates the identification of successfully modified BAC clones. This cassette is then removed by Flp-mediated recombination, leaving behind one FRT site.
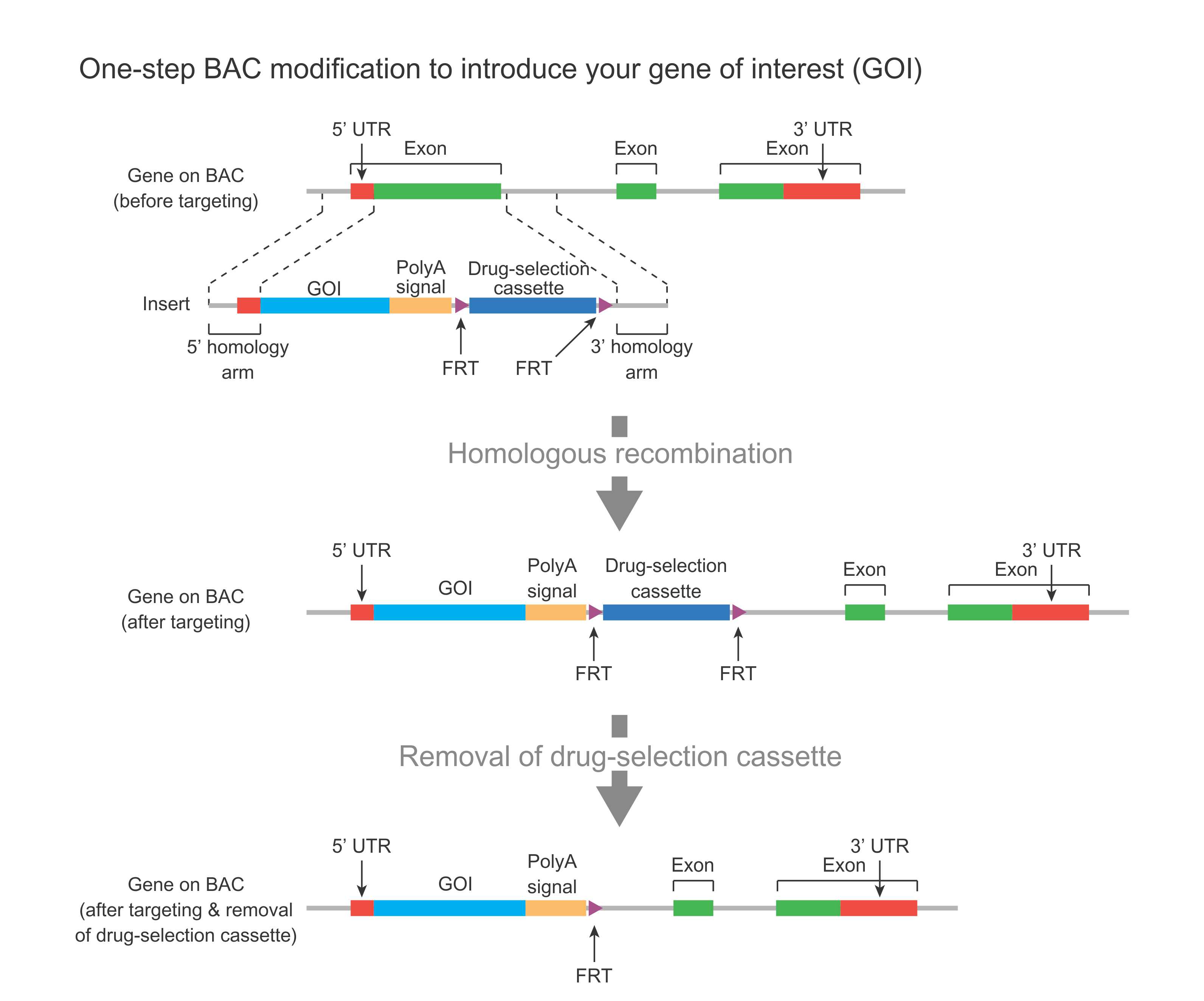
Figure 1. Schematic representation of one-step BAC modification for introducing a gene of interest (GOI) such as LacZ or GFP reporter behind the promoter of a gene.
In the case of a point mutation (Figure 2), a DNA fragment containing the mutated region, plus a drug-selection cassette flanked by FRT sites (along with flanking homology regions) is used to replace the corresponding endogenous region via homologous recombination. This cassette can then be removed by Flp-mediate recombination, leaving behind one FRT site. This leftover FRT is considered a “scar” sequence because the BAC now carries this leftover FRT in addition to the desired point mutation. In order not to affect gene function, the scar sequence should be placed in a nonfunctional region of the gene, such as an intron or UTR.
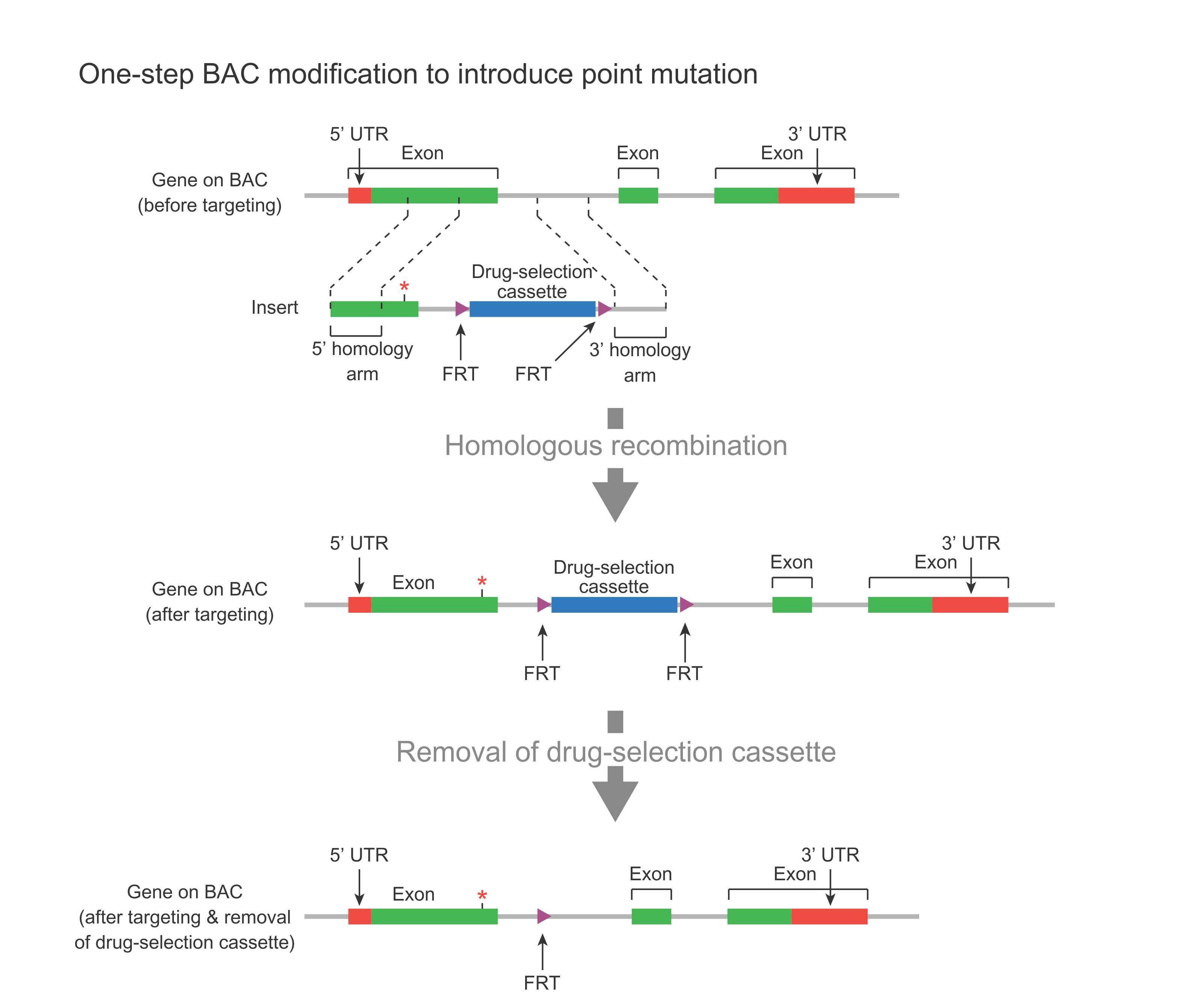
Figure 2. Schematic representation of one-step BAC modification for introducing a point mutation in the first exon of a gene along with an FRT scar sequence in the downstream intron.
Two-step BAC modification
The two-step modification method is used to introduce minor changes such as point mutations into a BAC without leaving behind the FRT scar sequence (Figure 3). This approach is preferred over the one-step approach when there is no place in the vicinity of the desired mutation for the leftover FRT scar sequence. One example where a two-step BAC modification is preferred is when the gene to be modified is a very large single-exon gene, and the desired point mutation is in the middle of the gene. Thus, there is no location near the point mutation to leave the scar sequence. The two-step method relies on positive and negative selection using a suitable drug-selection cassette (Figure 3). In the first step, this cassette is used to replace the target region via homologous recombination, which is facilitated by positive selection. In the second step, the cassette is further replaced with the final sequence containing the desired mutation via homologous recombination, which is facilitated by negative selection.
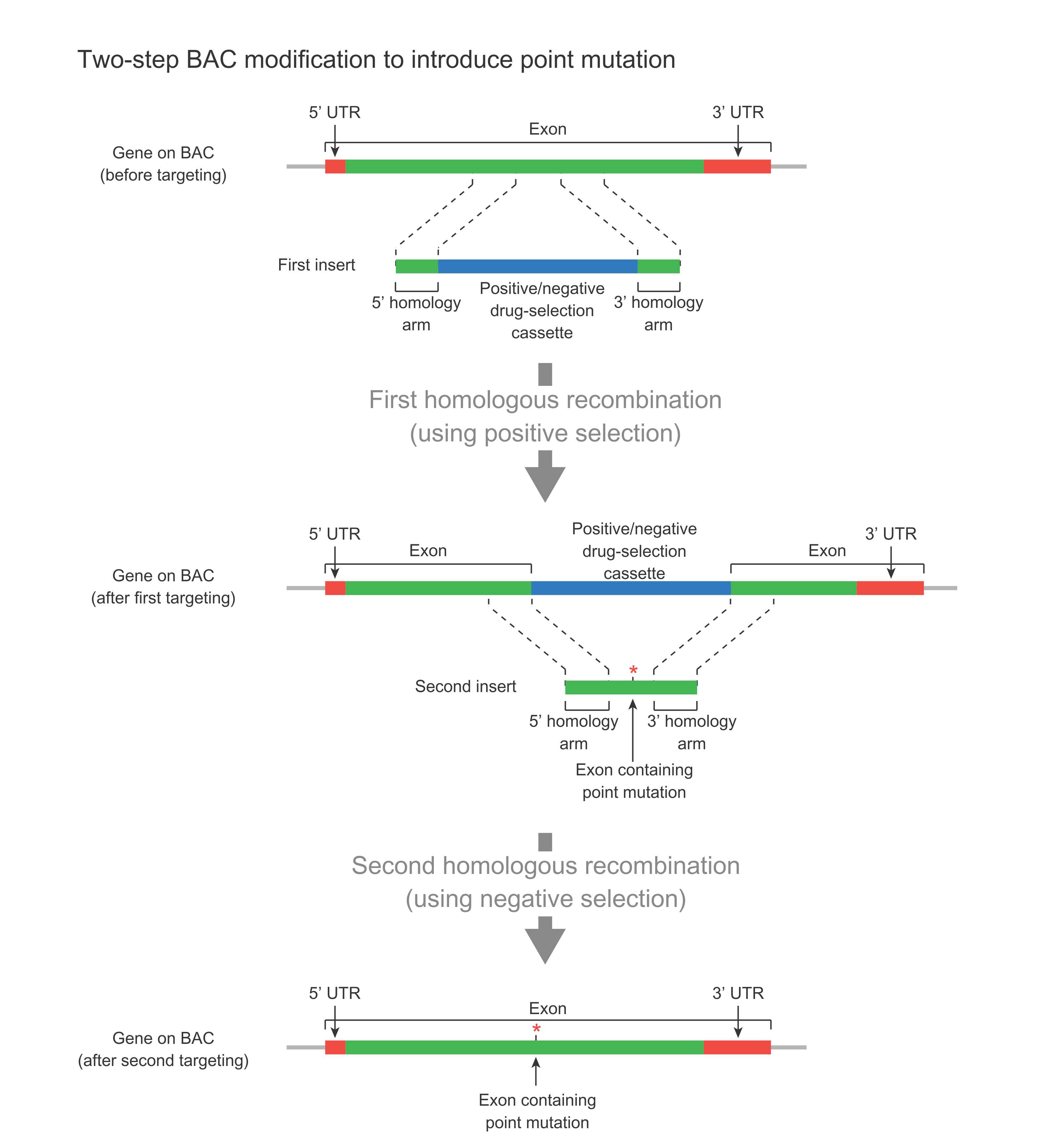
Figure 3. Schematic representation of two-step BAC modification for introducing a point mutation in a large single-exon gene without leaving behind any scar sequence.
Add drug-selection or visualization marker to BAC backbone
Adding a drug-selection marker such as neomycin resistance or a visualization marker such as EGFP to the BAC backbone facilitates the selection or visualization of cells successfully transfected with the BAC. This is especially useful for isolating cells containing stably integrated BACs by drug selection. Most BACs contain loxP sites on their backbones which can be utilized to insert a cassette expressing the marker of interest.
Transfer region of BAC onto plasmid
Transferring a specific region of a BAC onto a plasmid allows the region to be studied without the influence of flanking sequences. For example, if an entire BAC is used to create transgenic mice for studying the function of a gene on the BAC, the presence of other genes on the same BAC could confound the interpretation of the resulting phenotype. Isolating the gene onto a plasmid and making transgenic mice using the plasmid help circumvent this problem. An added benefit is the technical ease of working with plasmids versus BACs. We can transfer any region of a BAC (up to 60 kb) onto a plasmid.
How to Order
FAQ
What are the advantages of using BACs over YACs?
While both BACs and YACs are suitable for cloning large DNA fragments and even though YACs offer a larger cargo carrying capacity (up to 3000 Kb) compared to BACs (up to 300 Kb), BACs are typically preferred over YACs due to the following reasons: 1) BACs are more stable compared to YACs and therefore, help to maintain the integrity of large genomic DNA fragments. Using YACs for cloning large genomic DNA inserts often leads to the loss or rearrangements of DNA segments during propagation, thereby yielding erroneous end-products consisting of non-contiguous segments of the genome; 2) YACs exhibit significantly high levels (40-60 %) of chimerism caused by recombination between two YAC fragments or two YACs within the host yeast cells. Presence of such chimeric effects limits the use of YACs in various applications including contig assembly, chromosome walking and functional analysis of intact genes; 3) BACs can be propagated easily and rapidly in bacterial host cells and purified efficiently using standard DNA purification techniques like any other standard plasmid. YACs on the other hand are required to be propagated in and purified from yeast cells which is technically more challenging and time-consuming. Additionally, yeast cells typically yield considerably low amounts of YAC DNA upon purification.
Why are BACs preferred over standard expression plasmids for generating transgenic mice?
The ability of BACs to carry large DNA fragments allows large genomic DNA inserts including distant regulatory elements to be cloned into BACs. In contrast, standard expression plasmids can carry only the cDNA corresponding to the transgene of interest due to their limited cargo carrying capacity. The ability to include all regulatory elements associated with a given transgene within a BAC helps achieve optimal expression of the transgene, both spatially and temporally in transgenic animals harboring the BAC which is typically unachievable with a regular expression plasmid carrying only the transgene.




