Pooled shRNA Libraries
VectorBuilder offers high-quality premade pooled shRNA libraries targeting human and mouse genes. These RNAi libraries can serve as powerful and cost-efficient tools for large-scale loss-of-function screens across diverse biological processes. We can also construct custom shRNA libraries tailored to your research.
Types of pooled shRNA libraries offered
- Human gene targets
- Elite Gene (~2,000 most frequently cited genes on PubMed Central)
- Whole Genome (>20,000 RefSeq genes)
- Mouse gene targets
- Elite Gene (~2,000 most frequently cited genes on PubMed Central)
- Whole Genome (>20,000 RefSeq genes)
Ordering Information Price Match
| Product Name | No. of Genes | No. of shRNAs | Scale | Catalog No. | Price (USD) | Turnaround | Buy Now |
|---|---|---|---|---|---|---|---|
| Human Elite Gene Pooled shRNA Library | 2,161 | 12,471 |
Medium (>1.0x108 TU/ml, 1 ml) |
LVM(Lib190505-1037bjk) | $1,999 | 7-14 days | |
| Mouse Elite Gene Pooled shRNA Library | 2,233 | 12,472 |
Medium (>1.0x108 TU/ml, 1 ml) |
LVM(Lib190505-1039sgb) | $1,999 | ||
| Human Whole Genome Pooled shRNA Library | 20,593 | 105,233 |
Medium (>1.0x108 TU/ml, 1 ml) |
LVM(Lib230926-1079mym) | $1,999 | ||
|
Plus (>1.0x108 TU/ml, 5 ml) |
LV5M(Lib230926-1079mym) | $2,499 | |||||
| Mouse Whole Genome Pooled shRNA Library | 22,023 | 105,170 |
Medium (>1.0x108 TU/ml, 1 ml) |
LVM(Lib230926-1080rpt) | $1,999 | ||
|
Plus (>1.0x108 TU/ml, 5 ml) |
LV5M(Lib230926-1080rpt) | $2,499 |
Deconvolution service
Not sure how to identify shRNA hits by NGS after conducting your shRNA library screen? You can simply send the genomic DNA of your samples to VectorBuilder, and we will prepare the NGS libraries, perform high-throughput sequencing, analyze the data, and deliver the accurate counts of shRNAs in each sample to you in just a few weeks.
For deconvolution service, please Request Design Support from us.
Technical Information
Product highlights View more
Validation of library quality by NGS
VectorBuilder's libraries undergo rigorous validation through next-generation sequencing (NGS), and consistently demonstrate outstanding performance with high alignment scores and uniformity. The meticulous validation process ensures that the libraries are not only accurate but also maintain a high degree of consistency across sequences.
Genome-wide targeting and high complexity
For both human and mouse, we offer shRNA libraries at two scales: Whole Genome (~19,000 RefSeq genes) and Elite Gene (~2,000 most frequently cited genes on PubMed Central). On average, every gene is targeted by 5-6 different shRNAs with good knockdown scores calculated based on a set of rules following the guideline of the RNAi consortium (TRC). This enables more reliable knockdown and more effective screening. Detailed lists of target genes and shRNAs can be found under “Documents”.
High uniformity
The pooled plasmid libraries have been validated by next-generation sequencing (NGS), which has successfully detected 100% of shRNAs in the Elite Gene libraries and >97% of shRNAs in the Whole Genome libraries. In addition, shRNA representation is highly uniform in the plasmid libraries (Figure 1).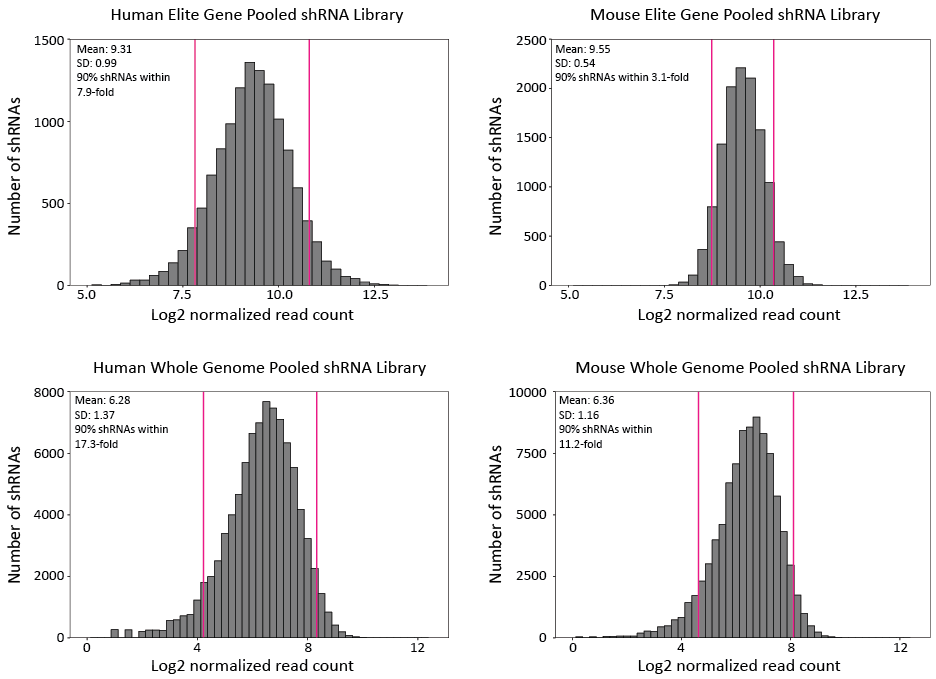
Figure 1. Representation of shRNAs in different pooled plasmid libraries. shRNA read counts are normalized by NGS library size (10 million reads) and plotted in log2 scale.
Well-established lentiviral vector and high titer lentivirus that is ready to use
The pooled shRNA libraries are expressed in the third-generation lentiviral vector system driven by human U6 promoter, which is a highly efficient system for stably knocking down expression of target genes in a wide variety of cells. This lentiviral vector system is ideal for in vitro genetic screens since it can introduce shRNAs into cells permanently, efficiently and uniformly. All pooled shRNA libraries are provided as ready-to-use lentivirus with high functional titer (>108 TU/ml), which saves your time in virus packaging and titer measurement. The third-generation lentiviral vector system is optimized for improved biosafety given its incompetent self-replication feature.
Figure 2. Map of mammalian shRNA knockdown lentiviral vector.
Click to view fully annotated map and sequence of the shRNA library vectorClick to read more about lentiviral shRNA knockdown vector
Dual-marker for efficient and versatile selection or tracking of positively transduced cells
A dual-marker expression cassette of EGFP and puromycin resistance gene (Puro) is expressed from the lentiviral vector, allowing for selection of positively-transduced cells by puromycin and visual tracking by green fluorescence.
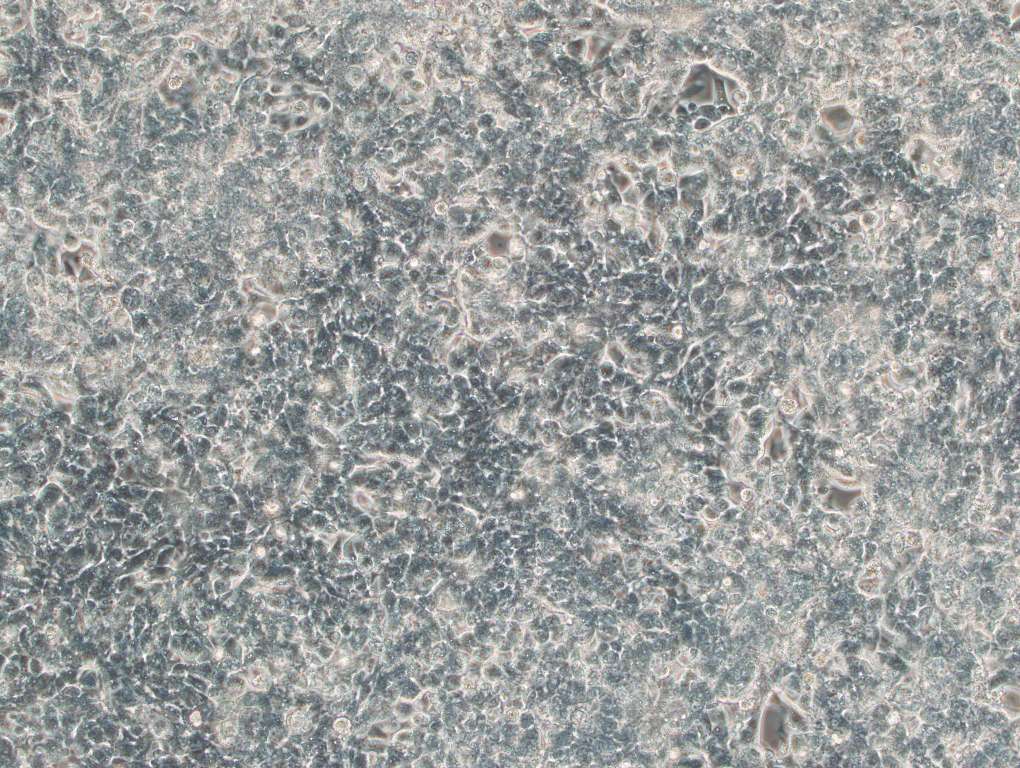

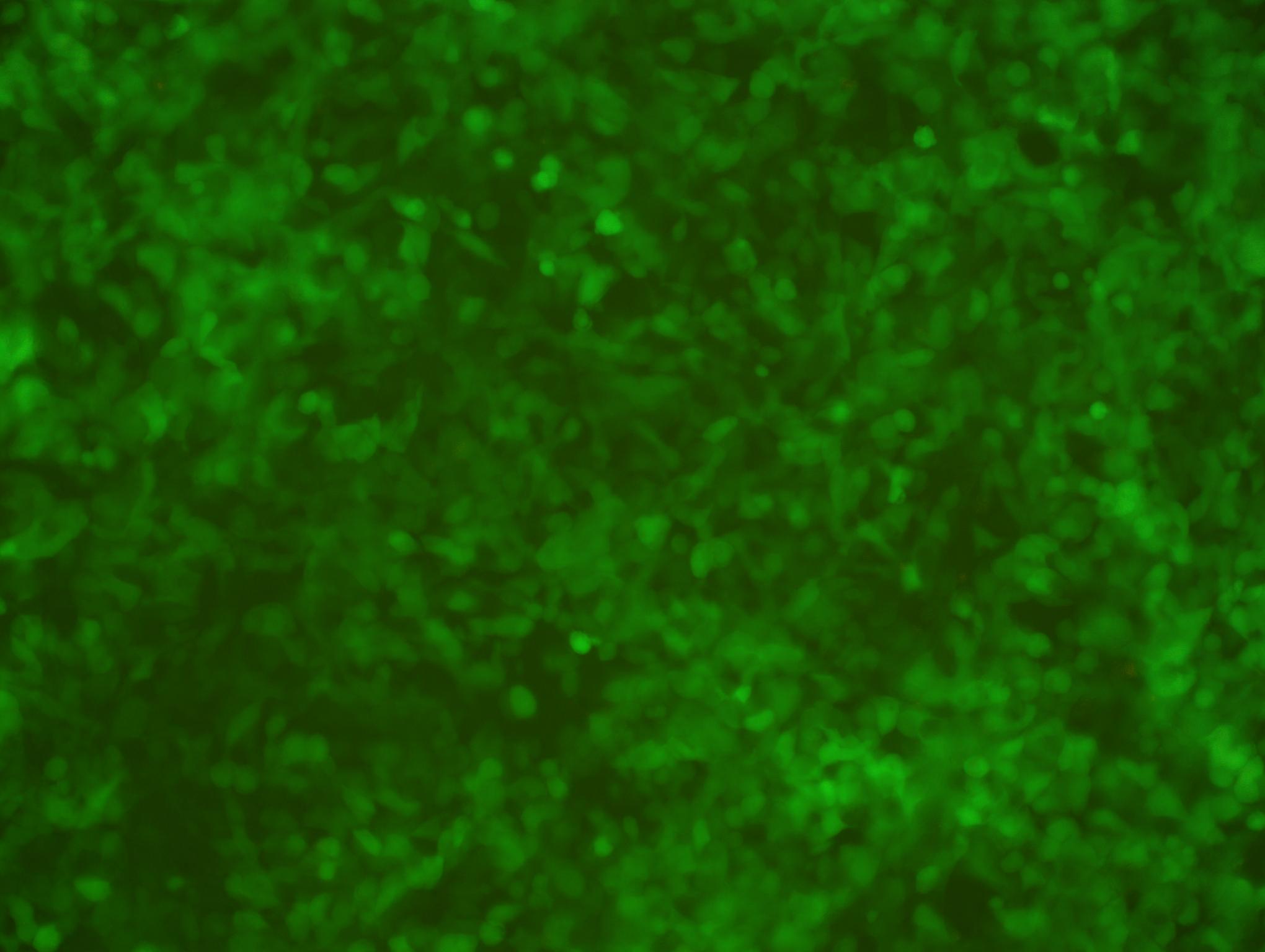
Figure 3. EGFP expression in 293T cells transduced with Human Elite Gene Pooled shRNA Library (MOI=10) after 4 days of puromycin selection (1.5 ug/ml). Magnification: 200x. Left: bright field. Right: EGFP.
Workflow of pooled shRNA library-mediated genetic screens View more
A conventional workflow of genetic screens mediated by pooled shRNA lentivirus libraries is shown in Figure 4 below. First, cells of interest are transduced with the shRNA lentivirus library, and positively transduced cells are selected by the marker gene(s) carried on the lentiviral genome (e.g. drug-selection or fluorescence marker). Next, positively transduced cells are split into reference population and experimental population. Then, the experimental population is subjected to particular selective pressure (e.g. drug treatment or repeated passaging) to identify cells with the phenotype(s) of interest. There are three major types of screening strategies: 1) viability screens that search for shRNAs enriched or depleted in surviving cells when exposed to selective pressure; 2) reporter screens that look for shRNAs enriched in cells associated with either high or low reporter expression (e.g. shRNAs targeting transcription factors that modulate reporter gene expression); 3) behavior screens that usually identify shRNAs affecting genes associated with cell invasion, migration, etc. After screening, both experimental and reference cells are harvested, and enriched or depleted shRNAs in the experimental group compared to the reference group are identified using Sanger sequencing or next-generation sequencing. Candidate genes that are potentially targeted by the enriched or depleted shRNAs can be further investigated by downstream functional studies.
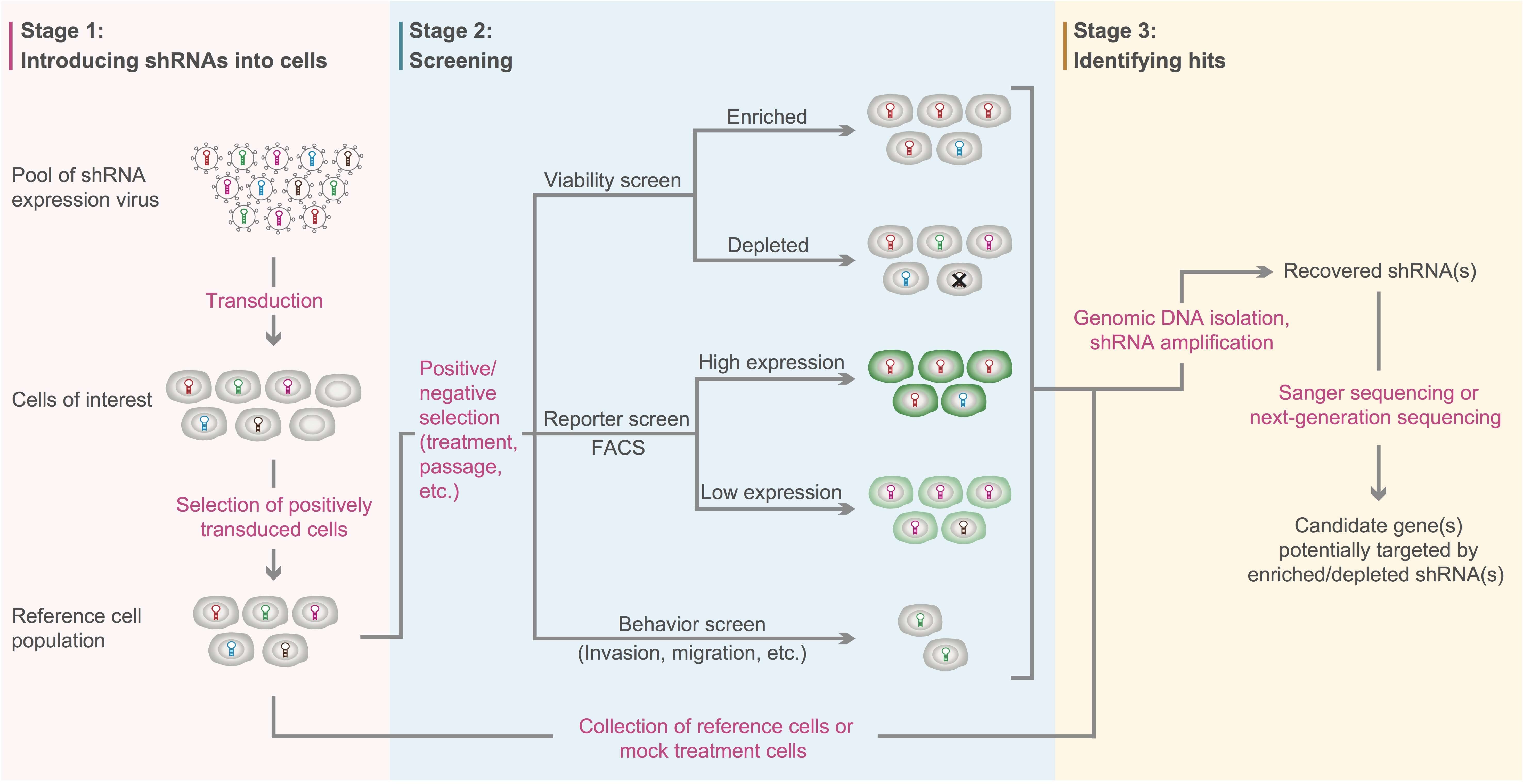
Figure 4. Workflow of loss-of-function screens mediated by pooled shRNA lentivirus libraries. Adapted from Acta Biochim Biophys Sin 44:103-112 (2012).
Resources
Documents
Full Lists of Target Genes and shRNAs
FAQ
What are the advantages of pooled RNAi screen compared to array-based RNAi screen?
Pooled RNAi screen has a number of advantages compared to array-based RNAi screen, which are listed in the table below:
| Array-based Screen | Pooled RNAi Screen | |
|---|---|---|
| How are shRNAs introduced into cells of interest? | Individual shRNAs are applied to cells grown in different wells across multi-well plates (e.g. 96-well or 384-well). | Hundreds and thousands of different shRNAs are applied to the cell population simultaneously. |
| How to identify shRNAs associated with phenotypes of interest? | Wells showing phenotypes of interest are selected by examining phenotypes well-by-well. The identity of the shRNAs applied to individual wells are already known. | Cells with phenotypes of interest are selected from the population. shRNAs enriched/depleted in cells of interest are identified by sequencing. |
| Can genetic interactions be detected? | No (if a single shRNA is added to each well), or limited (if more than one shRNAs are added to each well). | Yes (cells may carry multiple randomly combined shRNAs). |
| Technical variation | High | Low |
| Cost of labor and reagents | High | Low |
| Requirement of special equipment | High (e.g. liquid handler, high-throughput imaging, etc.) | Low (conventional benchtop equipment) |
How is shRNA knockdown score calculated?
VectorBuilder applies rules similar to that used by the RNAi consortium (TRC) to design and score shRNAs. For each given RefSeq transcript, we search for all possible 21mers that are considered as candidate target sites. Candidates are excluded if they contain features thought to reduce knockdown efficiency/specificity or cloneability, including a run of ≥4 of the same base, a run of ≥7 G or C, GC content <25% or >60%, and AA at the 5’ end. Knockdown scores are penalized for candidates that contain internal stem-loop, high GC content toward the 3’ end, known miRNA seed sequences, or off-target matches to other genes. For genes with alternative transcripts, target sites that exist in all transcripts are given higher scores.
All scores are ≥0, with mean at ~5, standard deviation at ~5, and 95% of scores ≤15. An shRNA with a knockdown score about 15 is considered to have the best knockdown performance and cloneability, while an shRNA with a knockdown score of 0 has the worst knockdown performance or is hard to be cloned.
Please note that knockdown scores are only a rough guide. Actual knockdown efficiency could depart significantly from what the scores predict. Target sites with low scores may still work well. Also, please note that targeting 3’ UTR can be as effective as targeting coding region.



