Premade IVT RNA and LNP-RNA
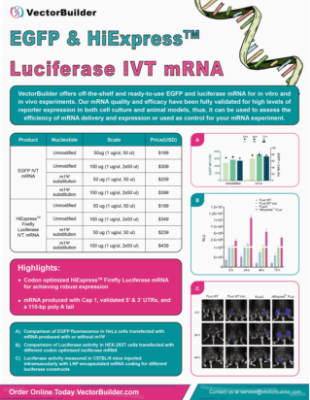
VectorBuilder offers ready-to-use IVT RNA and LNP encapsulated IVT RNA that has been fully validated in vitro and in vivo, thus, it can be used as control for your RNA experiment or to assess the efficiency of LNP-based RNA delivery systems. For custom IVT RNA and LNP production, check out VectorBuilder’s Therapeutic IVT RNA and LNP Encapsulation pages. For large-scale manufacturing of RNA drugs, check out our CDMO services.
Highlights
- IVT mRNA with Cap1, validated UTRs, uniform polyA tails, and the ability to add modified nucleotides, such as N1-Methylpseudouridine (m1Ψ)
- Premade circRNA and saRNA products for investigating next generation RNA therapeutics
- Sequence optimized HiExpressTM IVT RNA for achieving robust expression
- Comprehensive and strict quality controls to ensure identity, purity, and functionality
Product Information
Reporter gene mRNA
- EGFP IVT mRNA
- Zebrafish EGFP IVT mRNA
- mCherry IVT mRNA HOT
- HiExpress™ Firefly Luciferase IVT mRNA HOT
- HiExpress™ Gaussia Luciferase IVT mRNA
Antigen mRNA
CAR mRNA
Transposase mRNA
Reporter gene mRNA
- EGFP LNP-mRNA
- HiExpress™ Firefly Luciferase LNP-mRNA
- HiExpress™ Firefly Luciferase GalNAc LNP-mRNA NEW
Reporter gene saRNA
Reporter gene circRNA
Shipping and storage
Our RNA products are stored in a 1 mM sodium citrate buffer (pH 6.4) and can be stored at -80°C for up to 12 months. LNP-RNA products are stored in Tris buffer (pH 7.4) and can be stored at -80°C for up to 6 months. Both products are shipped on dry ice and repeated freeze-thaw cycles should be avoided.
Resources
Documents
Brochures & Flyers User InstructionsCertificate of Analysis (COA)
Material Safety Data Sheet (MSDS)
FAQ
What are the differences between mRNA caps and capping methods?
Cap 0 refers to N7-methylguanosine (m7G) that is added to the 5’ end of eukaryotic mRNAs via a 5’ to 5’ triphosphate linkage. This modification is added via a series of enzymatic reactions that occur co-transcriptionally and functions to regulate nuclear export, transcript stability, and promotes translation of the mRNA through recognition by eukaryotic translation initiation factor (eIF4E). Cap 1 refers to the addition of a methyl group to the 2’O on the first nucleotide (m7GpppNm) of the transcribed mRNA sequence in addition to the m7G cap. In mammalian cells, cap 1 structure is an important marker for mRNA to be recognized as self and not targeted by innate immunity. Adding cap 1 structure to synthesized mRNA has been demonstrated to enhance mRNA expression in vivo and reduced its immunogenicity.
Capping for in vitro transcribed RNA can occur either co-transcriptionally with cap analogs or post-transcriptionally via enzymatic reactions. The efficiency of our capping method has been well validated using LC-MS.
Why should I consider incorporating modified nucleotides in mRNA and which ones can be included?
Cells contain cytosolic and endosomal RNA receptors that activate the immune response upon recognition of foreign RNA. Modified nucleotides are commonly found in endogenous cellular RNA. Incorporating certain modified nucleotides in mRNA reduces its immunogenicity, alters secondary structure, and increases translation efficiency and half-life in a sequence-dependent manner. We provide the commonly used N1-Methylpseudouridine (m1Ψ). N1-Methylpseudouridine (m1Ψ) is a naturally occurring nucleotide that was first identified in tRNAs, however, its use in coding mRNAs has only recently been appreciated. This methylated derivative of uridine can replace the canonical nucleotide uridine in mRNA IVT and translation without altering traditional Watson-Crick base pairing. A major advantage of its use in mRNA therapeutics is its ability to alter recognition by RNA immune receptors thus mitigating unwanted immune effects and enhancing transcript stability and translation.