Snip Snap: Time for Transposons
When designing gene delivery experiments, one of the first questions to address is: will you use viral or non-viral delivery? While there are a huge number of viral vector options, it’s also easy to underestimate the options available for non-viral delivery. The second question researchers must ask is: do you need long-term or short-term gene expression? Long-term expression can be achieved with non-viral vectors using plasmids that carry transposons and transposases. But which system do you use? How do you use them? How can they be optimized? VectorBuilder is here with the answers to all your burning jumping gene questions.
Transposons: jump around (Jump! Jump!)
Even before the elucidation of how DNA is transcribed and translated into protein in the 1960’s, genetic elements were observed moving around chromosomes in plants. These transposable elements, or transposons, were later also found in everything from viruses to mammals. Within decades, the explosion of the field of molecular biology allowed for transposons to not only be studied but also edited and utilized for introduction of genetic material to host cell genomes.
In nature, DNA transposons exist as terminal repeats (TRs) flanking the coding region for a transposase. This transposase enzyme introduces double strand cuts at the TRs as well as genomic sites, allowing translocation of the entire transposon region. Continued expression of the transposase facilitates continued translocations. This system can easily be adapted for gene delivery by separating the transposons from the transposase and placing the genetic material to be inserted between the TRs (Figure 1).
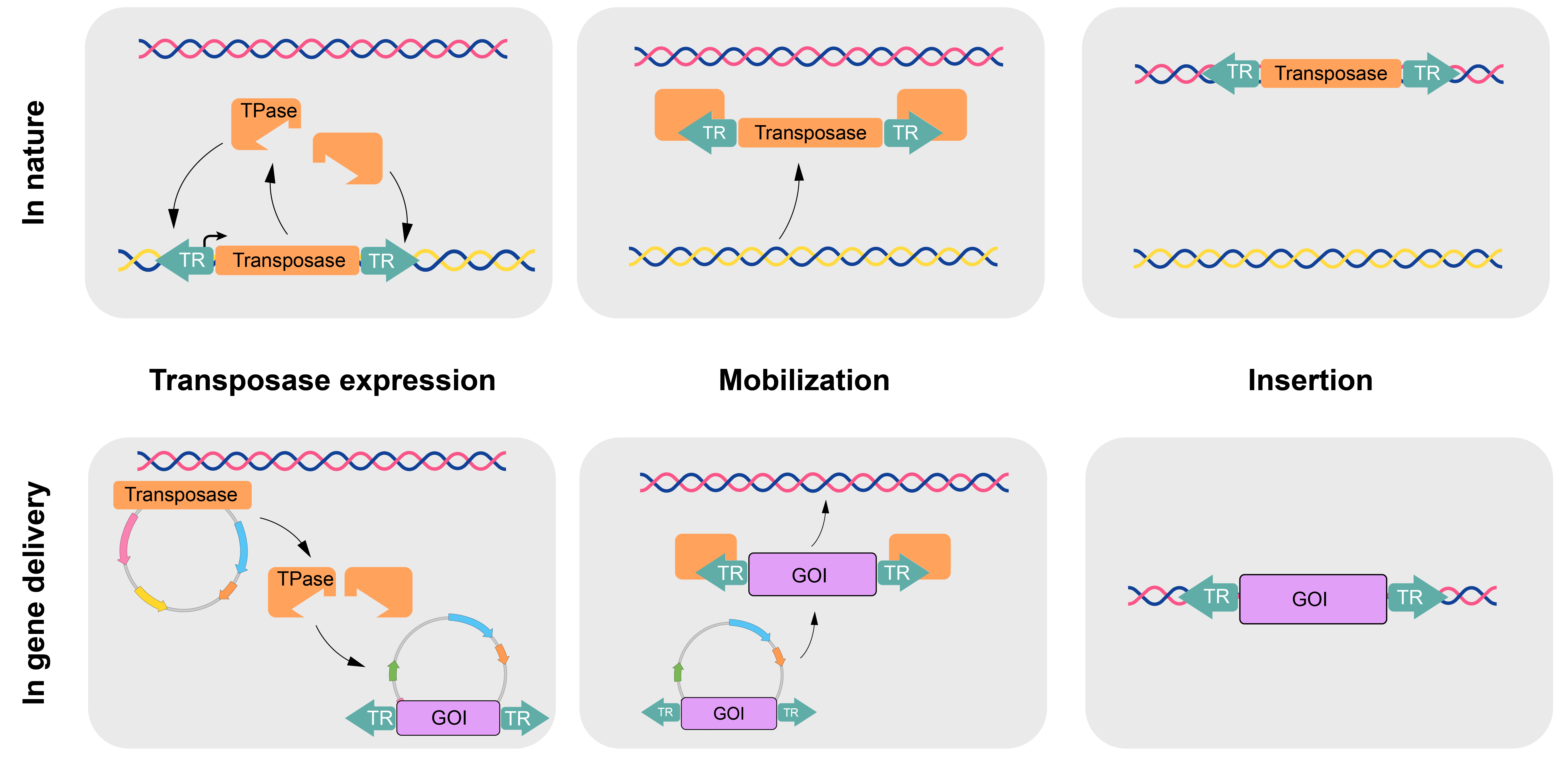
Figure 1. Transposons in nature can be separated into components on plasmids for gene delivery.
DNA transposons possess important advantages compared to viral counterparts like lentivirus: they have much lower cost of production and are more technically simple than viral vectors, and there is a lower risk of significant immune response. However, utilizing plasmids also has significant disadvantages, primarily difficulty in transfecting many cell types and limitations in in vivo transfection.
Plasmid picking
After deciding on a non-viral vector system with genomic integration, there are multiple systems to choose from: piggyBac, Sleeping Beauty, and Tol2. Each utilizes standard components:
1. A “transposon plasmid” containing the GOI (with related components including promoter, polyA tail, etc.) flanked by terminal repeats (TRs).
2. The corresponding transposase. This can be delivered as a “helper plasmid” or mRNA, or a cell/animal line stably expressing the transposase can be used.
Note: Only matching transposase and transposons can be used for efficient genomic integration, i.e., piggyBac transposase (PBase) cannot be used effectively with Sleeping Beauty or Tol2 transposons.
PiggyBac
One of the most popular transposon systems is the piggyBac vector system, which can be used to introduce a gene, shRNA, recombinant antibodies, and more. This system can also be combined with other approaches like CRISPR/Cas9 genome editing. The large carrying capacity of piggyBac’s transposon vector lends to its popularity, as it can carry up to 27 kb of DNA between the 5’ and 3’ TRs. The piggyBac transposase cuts chromosomal DNA at TTAA sites for insertion, with a preference for intragenic regions.
Since initial discovery, modifications have been made to both piggyBac transposons and transposase to increase efficiency: the transposons have been truncated, and the transposase has undergone codon optimization and mutagenesis leading to hyperactivity. While the native transposase PBase has higher efficiency than Sleeping Beauty or Tol2 transposases, the modified version hyPBase can achieve up to 15-fold higher transposition efficiency. Additionally, a further modified hyPBase is excision competent and integration incompetent, ensuring that integrated inserts are removed without reinsertion, leaving no footprint behind. Of note, hyPBase currently has an associated patent so should be used in development of commercial vectors only with appropriate counsel.
Use our Vector Design Studio to design your custom piggyBac vector with several promoters, ORFs, and markers to choose from.
Sleeping Beauty
The Sleeping Beauty (SB) vector system is also popular, particularly in clinical settings like CAR T cell therapy due to its tendency for random integration and reliance on a TA dinucleotide cut site. Similar to the piggyBac system, SB has an efficient transposase which has been modified into a hyperactive variant (SB100X). However, SB100X has lower efficiency in most cell types compared to hyPBase. Additionally, SB has a lower cargo capacity of about 8 kb and does leave a CAG footprint following excision.
While all three integrating vector systems have the significant restriction of limited target cell type range, Sleeping Beauty has been combined with viral vector systems including HSV and adenovirus to enhance cell targeting while maintaining integration capabilities.
Use our Vector Design Studio to design your custom Sleeping Beauty vector with several promoters, ORFs, and markers to choose from.
Tol2
Due to its lack of sequence bias and tendency to integrate into more flexible genomic regions, Tol2 is a commonly used system in the establishment of animal lines. The system's carrying capacity is moderately high, at 11 kb, and transposase activity is high, although it tends to be lower than piggyBac or SB transposases. Notably, Tol2 has high activity in fish models and is the historical go-to for creation of zebrafish lines.
Use our Vector Design Studio to design your custom Tol2 vector with several promoters, ORFs, and markers to choose from.
The table below lays out key comparisons between piggyBac, Sleeping Beauty, and Tol2 vector systems.
| Popular Transposases | Transposase Activity | Carrying Capacity | Integration Preference | Sequence Bias | |
|---|---|---|---|---|---|
| PiggyBac | PBase, hyPBase, PBx | Highest | 27 kb | Intragenic insertion | TTAA |
| Sleeping Beauty | SB, SB100X | High | 2-8 kb | Random insertion | TA |
| Tol2 | Tol2 | High | 11 kb | Intergenic insertion | No preference |
Transposon troubles
Like all adventures in gene delivery, setting up a transposon experiment requires careful planning and attention to controls. Because of the multiple components utilized and therefore multiple variables, negative controls are particularly important. For instance, for in vitro cultures, a negative control containing just the transposon vector should be set up in parallel to the experimental culture, and both should be sub-cultured until transgene expression is lost from the negative control. This indicates that the transgene only remains in cells in which transposition has occurred. Location of the transgene within the host genome can be confirmed with inverse PCR or linker-mediated PCR, which amplifies genomic DNA-transposon junctions.
When transposition efficiency is low, the problem may lie in the transposon vector, with the transposase, or both. The first and easiest step to check is the transposon to transposase ratio. For piggyBac systems using PBase, we recommend starting with a 1:1 ratio, with further details here. Increasing the transposase concentration may increase the integration efficiency, but it may also result in higher genotoxic effects. Instead, increasing the concentration of transposon vector will more likely improve efficiency. Additionally, efficiency can be increased by utilizing a more compact, miniaturized vector that forgoes bacterial backbone components. This not only enhances efficiency due to the smaller overall vector size but also avoids the risk of genomic integration of bacterial components.
How you introduce the transposase can also impact integration efficiency and stability. The two most popular options are transposase plasmid and mRNA. While the plasmid can have higher efficiency, there is the risk of longer-term transposase expression and activity as well as the low risk of genomic integration of the transposase. In both cases, continued activity of the transposase may lead to excision and reintegration of the transposon with potential host gene disruption. Utilizing transposase mRNA results in decreased efficiency but ensures short-term expression with no risk of transposase integration.
Conclusion
Non-viral vectors represent an important and ever-improving approach for areas of gene delivery, including studying gene function and delivering cell therapies. Optimizations are continuing for both transposons and transposases in the variety of options, so researchers can be sure that they are able to utilize the best possible approach for their study. Whether you are deciding which vector system, which transposase, or which insert to use, we’re here to help!
Source
de Silva S, Bowers WJ. Targeting the central nervous system with herpes simplex virus / Sleeping Beauty hybrid amplicon vectors. Curr Gene Ther. 2011 Oct;11(5):332-40. doi: 10.2174/156652311797415845. PMID: 21711226; PMCID: PMC4141986.
Eckermann KN, Ahmed HMM, KaramiNejadRanjbar M, Dippel S, Ogaugwu CE, Kitzmann P, Isah MD, Wimmer EA. Hyperactive piggyBac transposase improves transformation efficiency in diverse insect species. Insect Biochem Mol Biol. 2018 Jul;98:16-24. doi: 10.1016/j.ibmb.2018.04.001. Epub 2018 Apr 10. PMID: 29653176.
Kolacsek O, Erdei Z, Apáti A, Sándor S, Izsvák Z, Ivics Z, Sarkadi B, Orbán TI. Excision efficiency is not strongly coupled to transgenic rate: cell type-dependent transposition efficiency of sleeping beauty and piggyBac DNA transposons. Hum Gene Ther Methods. 2014 Aug;25(4):241-52. doi: 10.1089/hgtb.2013.149. PMID: 25045962; PMCID: PMC4142840.
Largaespada DA, Collier LS. Transposon-mediated mutagenesis in somatic cells: identification of transposon-genomic DNA junctions. Methods Mol Biol. 2008;435:95-108. doi: 10.1007/978-1-59745-232-8_7. PMID: 18370070; PMCID: PMC3517914.
Moretti A, Ponzo M, Nicolette CA, Tcherepanova IY, Biondi A, Magnani CF. The Past, Present, and Future of Non-Viral CAR T Cells. Front Immunol. 2022 Jun 9;13:867013. doi: 10.3389/fimmu.2022.867013. PMID: 35757746; PMCID: PMC9218214.
Sandoval-Villegas N, Nurieva W, Amberger M, Ivics Z. Contemporary Transposon Tools: A Review and Guide through Mechanisms and Applications of Sleeping Beauty, piggyBac and Tol2 for Genome Engineering. Int J Mol Sci. 2021 May 11;22(10):5084. doi: 10.3390/ijms22105084. PMID: 34064900; PMCID: PMC8151067



