Cell Line Models for Coronavirus Research
Cell line models have played an indispensable role in coronavirus research, including the isolation, propagation and genetic engineering of the virus, as well as studying mechanisms of cell entry and how cells respond to infection. Different coronavirus species may require different types of cells for their isolation, propagation and mechanistic studies, which can be challenging given the diversity of coronavirus.
Coronavirus constitutes a vast group of viruses that are extremely widespread in nature, infecting virtually all mammals and birds examined. Hundreds of coronavirus species have been characterized thus far. Of these, a few dozen can be deemed important because they infect humans, livestock, pets, or model animals, or they are evolutionarily close related to them. The phylogenetic tree of these important coronavirus species is shown below:
Phylogenetic Tree of Important Coronavirus Species
More detailed information of these important coronavirus species, including their known host receptors, is listed below:
List of important coronavirus species
VectorBuilder offers a variety of cell line models as described below for studying the above coronavirus species.
Types of cell lines offered
The cell lines used in coronavirus research can be broadly classified into three categories:
(1) Virus growth cell lines for isolating and propagating the virus.
(2) Virus packaging cell lines for packaging recombinant vectors into live virus.
(3) Virus assay cell lines for studying mechanisms of viral entry into host cells and other aspects of virus biology.
Click here to view experimental validation of our virus assay cell lines
As detailed below, VectorBuilder offers a rich panel of cell lines that meet all these needs.
Coronavirus growth cell lines
These cells are naturally susceptible to infection by the coronavirus species of interest, and can be used to isolate and propagate the virus. For initial isolation of live virus from infected biological samples such as sputum or infected tissues, cells are inoculated with the samples and cultured. Successful infection can lead to cytopathic effect (CPE) in many but not necessarily all cases. Viruses produced by the cells are then validated by electron microscopy and sequencing. The cells used to isolate a given type of virus are typically also used to propagate the virus indefinitely and amply it to large quantities. These cells can also be used to study virus biology including cell entry.
Our CoronaGrow™ cell line panel below is designed to facilitate the isolation and propagation of most coronavirus species.
| Cell Line Name | Description | Coronavirus Tested* | Catalog No. |
|---|---|---|---|
| CoronaGrow-VeroE6 | Subclonal line of Vero-E6 cells derived from adult African green monkey (Chlorocebus aethiops) kidney. It is susceptible to infection by many coronavirus species including SARS-CoV-2, and is the cell line of choice for the isolation and propagation of many viruses. | SARS-CoV-2, SARS-CoV, MERS-CoV, PanCoV-GD/P2S, BatCoV-WIV16, BatCoV-Rs4231, BatCoV-WIV1, BatCoV-RsSHC014, PEDV | CL0001 |
| CoronaGrow-Huh7 | Subclonal line of Huh7 cells derived from human liver cancer. It is susceptible to infection by many coronavirus species including HCoV-229E. | HCoV-229E, SARS-CoV-2, SARS-CoV, MERS-CoV | CL0002 |
| CoronaGrow-Frhk4 | Subclonal line of FRhK-4 cells derived from rhesus macaque (Macaca mulatta) fetal kidney. It is susceptible to infection by some coronavirus species including SARS-CoV. | SARS-CoV, CivetCoV-SZ3 | CL0003 |
| CoronaGrow-MK2 | Subclonal line of LLC-MK2 cells derived from rhesus macaque (Macaca mulatta) adult kidney. It is susceptible to infection by some coronavirus species including HCoV-NL63. | HCoV-NL63, SARS-CoV | CL0004 |
| CoronaGrow-L2 | Subclonal line of L2 cells derived from rat lung. It is susceptible to infection by some coronavirus species including MHV. | MHV, RCoV | CL0005 |
| CoronaGrow-HRT18G | Subclonal line of HRT-18G cells derived from human colon cancer. It is susceptible to infection by some coronavirus species including HCoV-OC43. | HCoV-OC43, BCoV, ECoV, CRCoV, RbCoV | CL0006 |
| CoronaGrow-Neuro2a | Subclonal line of Neruo-2a cells derived from mouse neuroblastoma. It is susceptible to infection by some coronavirus species including PHEV. | PHEV | CL0007 |
| CoronaGrow-ST | Subclonal line of ST cells derived from pig testis. It is susceptible to infection by some coronavirus species including TGEV. | TGEV, PRCoV, PDCoV | CL0008 |
| CoronaGrow-Fcwf4 | Subclonal line of Fcwf-4 cells derived from cat fetal macrophages. It is susceptible to infection by some coronavirus species including FIPV. | FIPV, FCoV | CL0009 |
| CoronaGrow-A72 | Subclonal line of A-72 cells derived from dog fibroblasts. It is susceptible to infection by some coronavirus species including CCoV. | CCoV | CL0010 |
* For viruses listed under each cell line, the cells have been shown to support their isolation and propagation. However, it does not preclude the possibility that the cells could support the growth of other coronaviruses species not listed.
Note: For coronavirus species colored in red, the cell lines that they are listed under are recommended for their isolation and propagation.
Coronavirus packaging cell lines
These cells are used to package live virus from recombinant vectors engineered to carry wildtype or mutant versions of the viral genome of interest.
To perform virus packaging, the vector DNA or its in vitro transcribed RNA is transfected into packaging cells. Once entering the cells, the DNA or RNA hijacks cellular machineries to produce live virus. Packaging cell lines are selected for their high transfection and packaging efficiency, but they may not support the continuous propagation of the virus because they may not express the appropriate receptors for virus binding and cell entry. Therefore, viruses packaged by our CoronaPack™ cells may need to be propagated using appropriate CoronaGrow™ cell lines. To do so, 6-24 hours after transfection, the packaging cells or the supernatant containing the initially packaged virus can be placed on the appropriate CoronaGrow™ cells, which in turn will support continuous propagation of the virus. Successful virus packaging and propagation could lead to CPE in CoronaGrow™ cells in many but not necessarily all cases.
Our CoronaPack™ cell lines below are designed to facilitate the packaging of essentially all coronavirus species.
| Cell Line Name | Description | Catalog No. |
|---|---|---|
| CoronaPack-BHK | Subclonal line of BHK-21 cells derived from baby hamster kidney. It is widely used for packaging coronavirus from recombinant vectors. | CL0011 |
| CoronaPack-293T | Subclonal line of 293T cells derived from human embryonic kidney. It can be used as an alternative cell line for packaging coronavirus from recombinant vectors. | CL0012 |
Coronavirus assay cell lines
A major application of cell-based assays in coronavirus research is to study the mechanisms of viral entry into host cells. This can be done using cell lines that naturally express cell-surface receptors recognized by viral S proteins (e.g. the CoronaGrow™ cells described above). An alternative is to use cell lines that do not naturally express the receptors but which are engineered to stably express receptors from exogenously introduced transgenes. This approach has the advantage of ensuring that the cells express high levels of the receptors of interest. Importantly, this approach also allows the expression of mutant receptors to reveal the function of specific receptor domains and amino acid residues in facilitating viral entry.
These receptor-expressing cell lines can be used to assay viral entry by naturally isolated coronavirus or artificially packaged coronavirus from recombinant vectors. Alternatively, the assay can be performed using recombinant lentivirus pseudotyped with the S proteins of interest. A major advantage of using pseudotyped lentivirus is its minimal biosafety requirements.
Click here to view our lentivirus pseudotyped with coronavirus spike (S) proteins
Our CoronaAssay™ cell line panel below is designed to stably express transgenes encoding various known coronavirus receptors including human ACE2, DPP4 and ANPEP, as well as mouse Ceacam1. We also offer cell lines that express the human TMPRSS2 transgene, either by itself or together with the receptor transgene. TMPRSS2 encodes a transmembrane serine protease implicated in facilitating cell entry of many coronavirus species that utilize ACE2, DPP4, ANPEP or sialic acid as receptors. The CoronaAssay™ panel comprises subclonal lines from either HeLa (derived from human cervical cancer) or 293T (derived from human embryonic kidney) parental cells.
Click here to view experimental validation
| Cell Line Name | Transgene | Utility | Catalog No. | |
|---|---|---|---|---|
| 293T-Based Cell Lines** | CoronaAssay-293T | None | No-transgene control | CL0012 |
| CoronaAssay-293T(hTMPRSS2) | Human TMPRSS2 | TMPRSS2-alone control | CL0013 | |
| CoronaAssay-293T(hACE2) | Human ACE2 | Assay coronavirus that targets ACE2 receptor (e.g. SARS-CoV-2) | CL0014 | |
| CoronaAssay-293T(hACE2-hTMPRSS2) | Human ACE2 Human TMPRSS2 |
Assay coronavirus that targets ACE2 receptor (e.g. SARS-CoV-2) | CL0015 | |
| CoronaAssay-293T(hACE2-hTMPRSS2-hNRP1) | Human ACE2 Human TMPRSS2 Human NRP1 |
Assay coronavirus that targets ACE2 receptor (e.g. SARS-CoV-2) | CL0016 | |
| CoronaAssay-293T(hANPEP) | Human ANPEP | Assay coronavirus that targets ANPEP receptor (e.g. HCoV-229E) | CL0017 | |
| CoronaAssay-293T(hANPEP-hTMPRSS2) | Human ANPEP Human TMPRSS2 |
Assay coronavirus that targets ANPEP receptor (e.g. HCoV-229E) | CL0018 | |
| HeLa-Based Cell Lines* | CoronaAssay-HeLa | None | No-transgene control | CL0019 |
| CoronaAssay-HeLa(hTMPRSS2) | Human TMPRSS2 | TMPRSS2-alone control | CL0020 | |
| CoronaAssay-HeLa(hACE2) | Human ACE2 | Assay coronavirus that targets ACE2 receptor (e.g. SARS-CoV-2) | CL0021 | |
| CoronaAssay-HeLa(hACE2-hTMPRSS2) | Human ACE2 Human TMPRSS2 |
Assay coronavirus that targets ACE2 receptor (e.g. SARS-CoV-2) | CL0022 | |
| CoronaAssay-HeLa(hACE2-hTMPRSS2-hNRP1) | Human ACE2 Human TMPRSS2 Human NRP1 |
Assay coronavirus that targets ACE2 receptor (e.g. SARS-CoV-2) | CL0023 | |
| CoronaAssay-HeLa(hDPP4) | Human DPP4 | Assay coronavirus that targets DPP4 receptor (e.g. MERS-CoV) | CL0024 | |
| CoronaAssay-HeLa(hDPP4-hTMPRSS2) | Human DPP4 Human TMPRSS2 |
Assay coronavirus that targets DPP4 receptor (e.g. MERS-CoV) | CL0025 | |
| CoronaAssay-HeLa(mCeacam1) | Mouse Ceacam1 | Assay coronavirus that targets Ceacam1 receptor (e.g. MHV) | CL0026 |
* HeLa-based cell lines have zero expression of endogenous ACE2, DPP4, CEAMCAM1 and TMPRSS2.
** 293T-based cell lines have zero expression of endogenous ANPEP and TMPRSS2
Technical documents
User InstructionsExperimental Validation
We developed a number of proprietary techniques to optimize our virus assay cell lines. As a result, our virus assay cell lines can be transduced by our optimized pseudotyped lentivirus with much higher efficiency as compared to published transducing protocols. We carried out extensive experimental validation of our transduced cell lines. The results showed that SARS-CoV-2 S protein and SARS-CoV-2 S protein with D614G mutation pseudotyped lentivirus could efficiently transduce 293T cells expressing high levels of the human ACE2 receptor, but not 293T cells in which ACE2 expression is absent or low, using fluorescent reporter (Figure 1) and luciferase reporter (Figure 2). We, therefore, recommend using our ACE2-expressing cell lines that have been optimized for transduction with SARS-CoV-2 S protein or VSV-G pseudotyped virus.
Click here to view our lentivirus pseudotyped with coronavirus S proteins
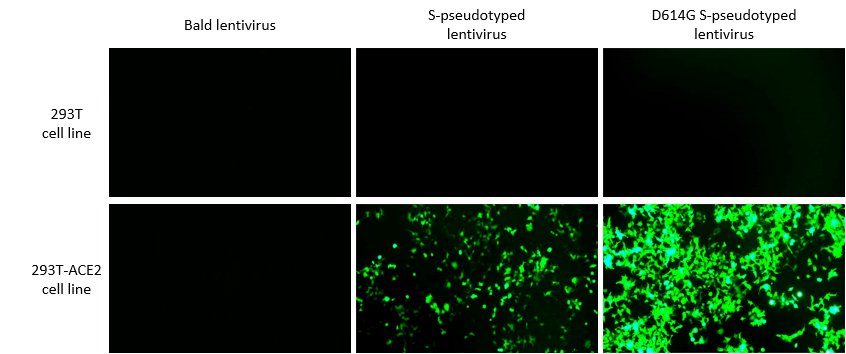
Figure 1. Lentivirus pseudotyped with SARS-CoV-2 S protein specifically infected 293T cells overexpressing human ACE2 receptor. Images were taken at 72 hours post-transduction.
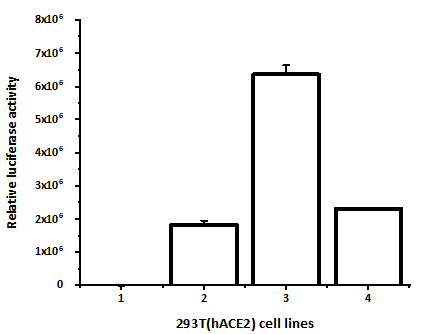
Figure 2. 293T(hACE2) cells transduced with lentivirus pseudotyped by different proteins. Column 1: Untransduced cells. Column 2: SARS-CoV-2 S protein. Column 3: SARS-CoV-2 D614G S protein. Column 4: VSV-G Protein.
How do I order cell lines?
You can inquire about our cell lines by following the link below:



