Precision Genetics: Navigating the Cre-lox Toolbox
Keywords: Cre recombinase, conditional knockout, Cre-lox system
The Cre-Lox system is a tool in genetic engineering that allows for precise control over gene expression and genetic modification. The system consists of two main components: a Cre recombinase and loxP sites. In this article we will review these two main components and delve into how they are applied in the lab every day. After an overview of the system, we will dive into advances within the system itself and how the system it is being used to advance discovery.
“Causes recombination” – “locus of crossover(x) P1”
Cre recombinase is an enzyme derived from the P1 bacteriophage, and it plays a central role in the Cre-lox system. The acronym Cre was initially derived from the term “causes recombination”; however, today it is also referred to by the shortening of “cyclization recombinase”. This enzyme possesses the unique ability to recognize and interact with specific DNA sequences known as loxP sites.
LoxP sites, short for “locus of crossover (x) P1”, are specific DNA sequences that consist of two 13 base-pair inverted and palindromic repeats separated by an 8 base-pair core sequence spacer region. Cre is a site-specific recombinase that recognizes the 13 base-pair inverted repeat with the sequence of the core region giving directionality to the Cre-lox system. When two loxP sites are placed facing opposite directions, the Cre recombinase inverts the segment of DNA between the two sites. Conversely, when two loxP sites flank a segment of DNA in the same direction, the Cre recombinase excises the segment between the sites resulting in deletion of genetic information. Pairing loxP sites facing the same direction in different locations in the genome allows for translocation (Figure 1).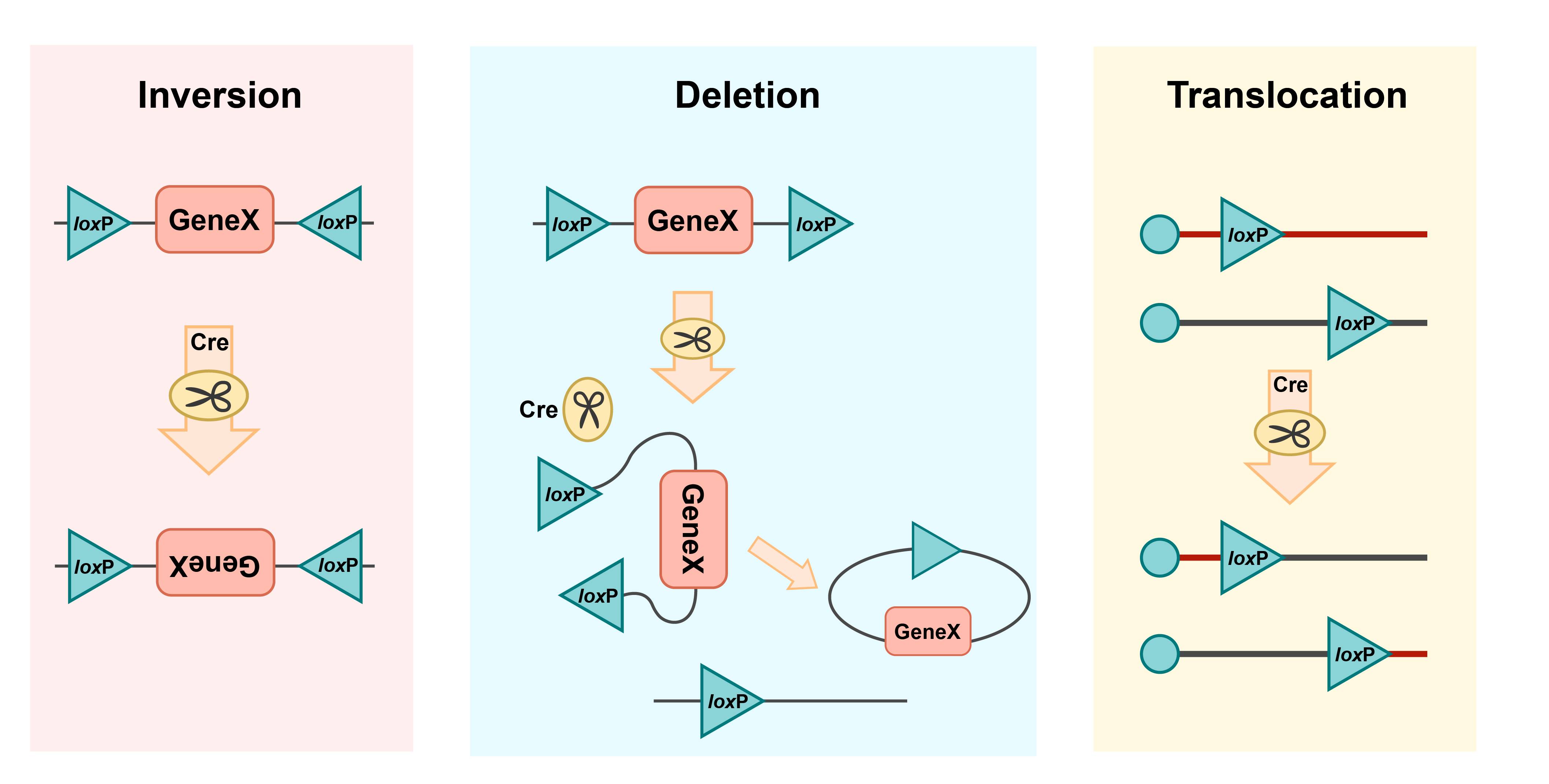
Figure 1. Mechanism of recombination with the Cre-lox system.
The unique properties of the Cre-lox system allow for its applications to several techniques in genetic engineering. One of the most popular applications of the Cre-lox system is its use in producing transgenic mice. In mice, the Cre-lox system is extensively employed as a tool for conditional gene manipulation. Researchers engineer mice with loxP sites strategically placed around a target gene, creating a “floxed” gene. These mice are then bred with transgenic mice expressing a Cre recombinase under the control of specific promoters, allowing spatial and temporal control of gene modification (Figure 2). When Cre is activated in specific tissues or at certain developmental stages, it catalyzes recombination at the loxP sites, leading to the precise alteration of the target gene. This enables conditional gene knockout, activation, or other modifications, offering insights into gene function in a controlled and dynamic manner. The Cre-lox system in mice has proven invaluable in understanding gene function, modeling diseases, and developing potential therapeutic interventions with a high level of precision.
For instance, conditional knockout models of the tumor suppressor gene p53 enable researchers to explore its role in maintaining genomic stability and preventing uncontrolled cell growth. Activation of oncogenes like KRAS through Cre-lox technology allows for the study of their specific contributions to tumor initiation and progression. Conditional knockout models of genes such as PTEN, BRCA1/BRCA2, and E-cadherin provide insights into their roles in various cancer types, including breast, ovarian, and colorectal cancers. These models, by allowing the targeted manipulation of specific genes in a tissue-specific or temporally controlled manner, serve as invaluable tools for unraveling the intricate molecular mechanisms underlying cancer development and for testing novel therapeutic interventions in a preclinical context.
In addition to utilization in mouse lines, the Cre-lox system is often employed in vitro for controlled genetic manipulations. This approach allows researchers to utilize non-viral vectors, such as plasmids or DNA constructs containing loxP sites to be easily introduced into cultured cells or cell lines. These loxP-flanked sequences can be engineered to carry genes of interest, reporter genes, or other functional elements. Subsequently, the cells can be transfected with a separate construct expressing Cre recombinase. The presence of Cre within the cells leads to site-specific recombination between the loxP sites, resulting in the desired genetic modification.
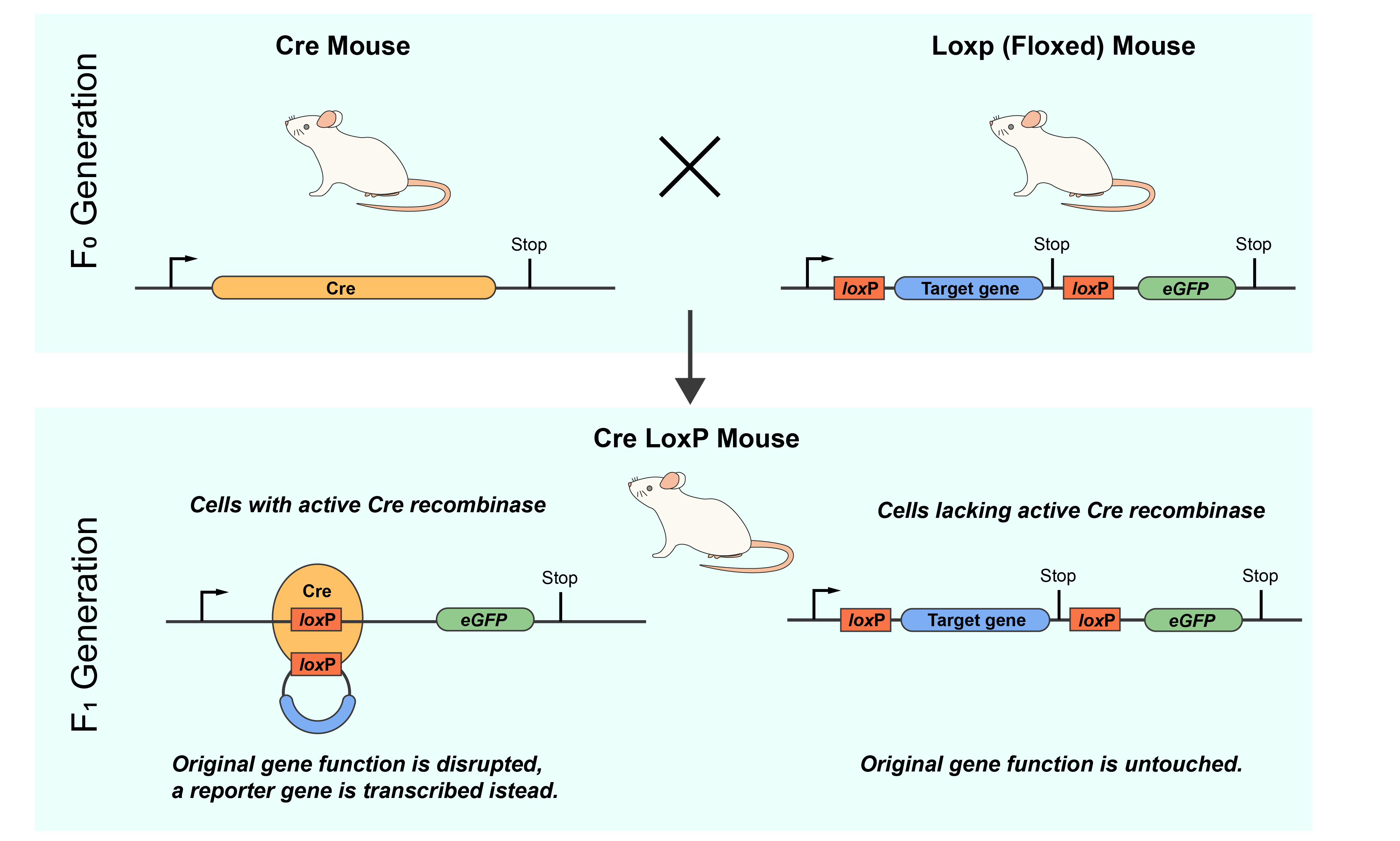 Figure 2. Mechanism of generation of tissue specific knockout mice using the Cre-lox system.
Figure 2. Mechanism of generation of tissue specific knockout mice using the Cre-lox system.
FLEX-ing a little
The Cre-lox FLEX system, also known as the FLEX switch, is an extension of the classical Cre-lox system designed to provide increased flexibility and control over genetic modifications. The term "FLEX" stands for "FLip-EXcision". In the Cre-lox FLEX system, the Cre recombinase is used in conjunction with modified loxP sites called "FLEX" sites. These sites are engineered to have asymmetric orientation and additional flanking sequences, allowing for bidirectional and reversible genetic modifications. The FLEX system is particularly useful for achieving temporal control over gene expression. When Cre recombinase is present, it can catalyze recombination between the FLEX sites, leading to a change in the orientation of the DNA segment between them. This change in orientation can result in either the activation or inactivation of a gene, depending on its initial configuration.
The "FLEX on" and "FLEX off" configurations refer to the states of a genetic construct designed using the Cre-lox FLEX system. In the "FLEX on" state, the genetic element of interest, such as a gene or reporter, is introduced in the reverse or antisense orientation, so transcription does not occur. However, it is activated or expressed because of inversion of the FLEX sites after Cre-mediated recombination. Conversely, in the "FLEX off" state, the same genetic element is introduced in a state of active expression but, upon introduction of Cre, is either inactivated or not expressed, as the new orientation of the FLEX sites prevents its transcription. The bidirectional and reversible nature of the Cre-lox FLEX system allows researchers to precisely control the activation or silencing of genes in a temporal and spatial manner, providing a powerful tool for conditional and dynamic gene expression studies. Cre-On vectors, often referred to as "DIO" vectors, signify the gene of interest being "Double-floxed with Inverted Orientation," while Cre-Off vectors, known as "DO" vectors, indicate the gene of interest being "Double-floxed in Orientation" (Figure 3). FLEX switches serve as research tools applicable to various scientific purposes, providing researchers with a powerful means to precisely regulate gene expression in a irreversible manner.
Another popular variation of this system deemed Cre-Switch allows for permanent Cre-mediated switching between the expression of two ORFs in mammalian in vitro and in vivo systems. The FLEX Cre-Switch system employs two pairs of loxP-variant recombination sites flanking two antiparallel ORFs, allowing activation of one gene and inactivation of the other through Cre-dependent inversion of both ORFs. The vector is designed with heterotypic loxP variants (loxP and lox2272), ensuring recombination specificity. Both loxP variants are recognized by Cre, but only identical pairs of loxP and lox2272 sites can recombine with each other and not with any other variant. In the absence of Cre, the first ORF is expressed under the user-selected promoter, while the second ORF remains silent due to its antisense orientation. In the presence of Cre, recombination events lead to the inversion of both ORFs, resulting in the silencing of the first and expression of the second.
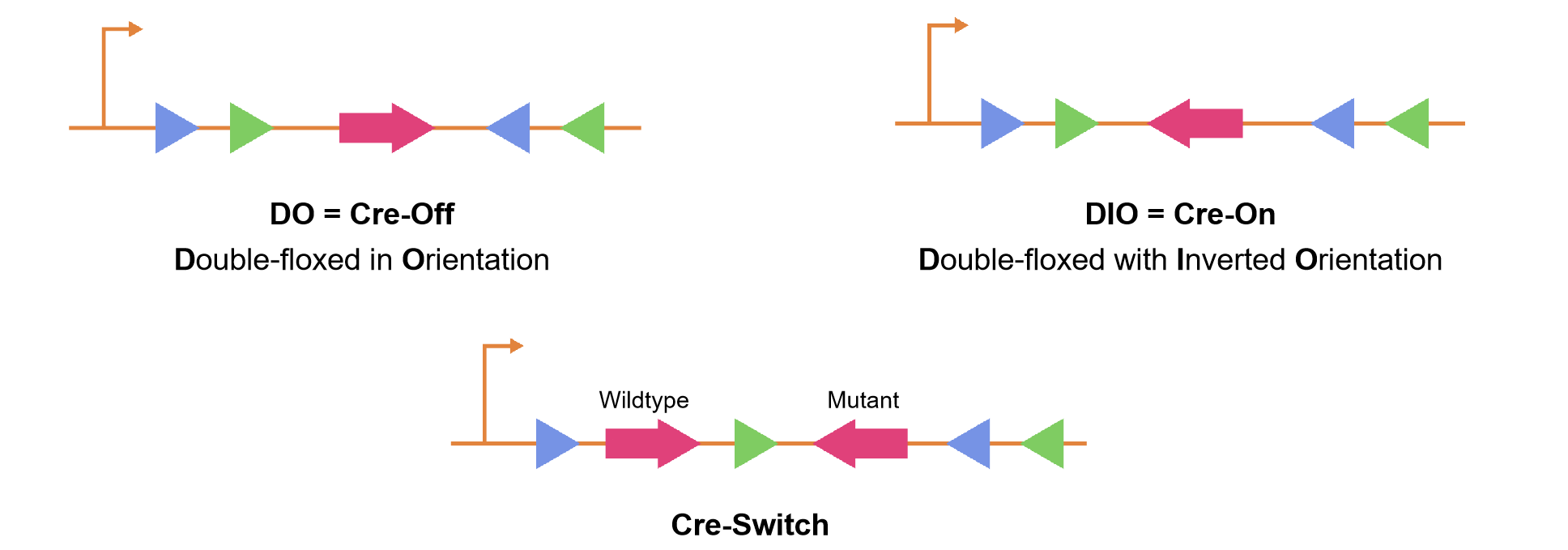
Figure 3. Diagram of gene and loxp site positioning in the Cre-On, Cre-Off, and Cre-switch.
Delivering Cre with AAV
Cre recombinase can be selectively delivered to specific cells using Adeno-Associated Virus (AAV) vectors driven by tissue-specific promoters. AAV vectors are widely employed due to their safety profile and efficiency in transducing various cell types. By incorporating tissue-specific promoters into the AAV vector, the expression of Cre recombinase can become tightly regulated, ensuring its activation only in specific cell types or tissues of interest. This precision allows researchers to control Cre recombinase activity in a spatially and temporally restricted manner. Consequently, when these AAV vectors are administered to experimental subjects, such as transgenic mice, the tissue-specific promoters guide the delivery of Cre recombinase to the desired cells, facilitating targeted genetic modifications. This approach is instrumental in studies requiring conditional gene manipulation, enabling researchers to investigate gene function in a highly specific and controlled fashion within cell populations or tissues. Conditional systems can also have enhanced temporal control through combination of Cre-lox and Tet inducible systems, where Cre recombinase expression can be controlled by tetracycline regulated promoters.
Recently, Cre-lox technology has been applied to study the role of TGF-B in tumor associated macrophages. TGFBI flox/flox mice were generated, allowing in vivo depletion of the TGFBI gene in specific cells or tissues. A Cre-recombinant virus (AAV8-mCD68-Cre) was constructed for cell-specific gene manipulation, with CD68 promoter ensuring Cre expression specifically in macrophages. TGFBI flox/flox mice received AAV8-mCD68-Cre virus, leading to TGFBI knockout in MΦs. The results showed significantly reduced tumor weights in TGFBI flox/flox mice with AAV8-mCD68-Cre injection compared to control groups, suggesting that TGFBI plays a role the tumor associated macrophage contribution pancreatic cancer tumor growth.
In conclusion, the use of AAV as a vehicle for delivering Cre recombinase serves as a powerful tool in genetic engineering. The precision and efficiency achieved through AAV-mediated delivery, especially when coupled with tissue-specific promoters, offer an unparalleled level of control in manipulating gene expression. This technology, as demonstrated in studies targeting TGFBI in macrophages for pancreatic cancer suppression, showcases the power of AAV in facilitating highly targeted and specific genetic modifications. The ability to utilize AAV vectors for Cre delivery not only enhances the accuracy of gene manipulation but also opens new avenues for investigating complex biological processes, developing innovative therapeutic strategies, and advancing our understanding of diseases at the molecular level.
Sources
Zhou, Jing, et al. "A novel role of TGFBI in macrophage polarization and macrophage-induced pancreatic cancer growth and therapeutic resistance." Cancer letters 578 (2023): 216457.
Schnütgen F, Doerflinger N, Calléja C, Wendling O, Chambon P, Ghyselinck NB. A directional strategy for monitoring Cre-mediated recombination at the cellular level in the mouse. Nat Biotechnol. 2003 May;21(5):562-5. doi: 10.1038/nbt811. Epub 2003 Mar 31. PMID: 12665802.
Lee G, Saito I. Role of nucleotide sequences of loxP spacer region in Cre-mediated recombination. Gene. 1998 Aug 17;216(1):55-65. doi: 10.1016/s0378-1119(98)00325-4. PMID: 9714735.



